2025 AIChE Annual Meeting
(400ao) Can Coarse-Grained Molecular Dynamics Simulations Predict Pharmaceutical Crystal Growth?
Author
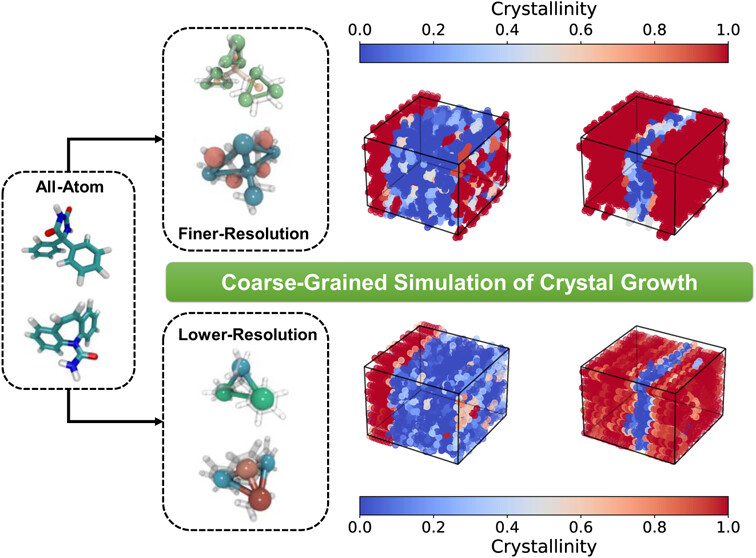
To investigate the ability of coarse-grained molecular dynamics simulations to predict the relative growth rates of crystal facets of pharmaceutical molecules, we apply two coarse-graining strategies to two drug molecules, phenytoin and carbamazepine. In the first method, we map an atomistic model to a MARTINI-level coarse-grained (CG) force field that uses 2 or 3 heavy atoms per bead. This is followed by applying Particle Swarm Optimization (PSO), a global optimum searching algorithm, to the CG Lennard-Jones intermolecular potentials to fit the radial distribution functions of both the crystalline and melt structures. In the second, a coarser-grained method, we map 5 or more heavy atoms into one bead with the help of the Iterative Boltzmann Inversion (IBI) method to derive a tabulated longer-range force field (FF). Simulations using the FF’s derived from both strategies were able to stabilize the crystal in the correct structure and to predict crystal growth from the melt with modest computational resources. We evaluate the advantages and limitations of both methods and compare the relative growth rates of various facets of both drug crystals with those predicted by the Bravais–Friedel–Donnay–Harker (BFDH) and attachment energy (AE) theories. While all methods, except for the simulations conducted with the coarser-grained IBI-generated model, produced similarly good results for phenytoin, the finer-grained PSO-generated FF using MARTINI mapping rules outperformed the other methods in its prediction of the facet growth rates and resulting crystalline morphology for carbamazepine.